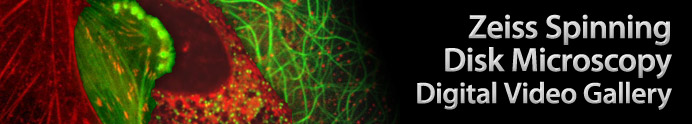
Monkey Kidney Cells with mEmerald-PMP and mRuby-ER
Peroxisomes possess an array of enzymes, which principally function together to clear the cell of toxic substances, especially hydrogen peroxide - a common byproduct of cellular metabolism. These organelles hold enzymes that convert the hydrogen peroxide to water, making the toxic substance safe enough to be discharged into the cell.
At any given moment, there exists a wide variety of biochemical compounds in the endoplasmic reticulum. For most, the organelle is a place of transitory habitation. After being formed and processed, the compounds are sent to other cellular locations. A few resident proteins, however, stay put in the lumen of the ER permanently. A specific amino acid sequence contained in resident proteins serves as a retention signal that lets the organelle know not to transport the proteins out of the ER. A prime example of an ER resident protein is BiP, a molecular chaperone that helps prevent aberrant proteins from reaching other parts of the cell.
The endoplasmic reticulum was visualized in the African green monkey kidney fibroblast cells (CV-1 line) featured in this digital video sequence with mRuby, a monomeric derivative of the tetrameric far-red fluorescent protein eqFP611. Like eqFP611, mRuby displays a large Stokes shift, but the newer fluorescent protein has the advantage of increased brightness. Excitation and emission maxima of mRuby occur at 558 and 605 nanometers, respectively. The green fluorescent protein employed in this section to visualize peroxisomes, mEmerald, is an EGFP variant that exhibits peak excitation at 487 nanometers and peak emission at 509 nanometers.